2.5.4
DBSCAN
- DBSCAN (Density-Based Spatial Clustering of Applications with Noise) groups points according to local density so clusters can take any shape while sparse regions become noise.
- Two hyperparameters control the model:
eps, the neighbourhood radius, andmin_samples, the minimum number of neighbours required for a point to become a core sample. - Points are labelled core, border, or noise; clusters are connected components of core points plus their border neighbours.
- A common tuning recipe is to fix
min_samples(≥ dimensionality + 1) and sweepepswhile inspecting the share of points flagged as noise.
Intuition #
This method should be interpreted through its assumptions, data conditions, and how parameter choices affect generalization.
Detailed Explanation #
1. Overview #
DBSCAN does not require the number of clusters beforehand. Instead, it inspects each sample:
- Core points: at least
min_samplesneighbours within distanceeps. - Border points: lie within the
eps-ball of a core point but fail the core criterion themselves. - Noise points: belong to no core neighbourhood.
Because of this density view, DBSCAN is robust to non-convex clusters such as two moons or concentric circles. Always scale the features so eps has a meaningful interpretation.
2. Formal definition #
Given (x_i \in \mathcal{X}), its (\varepsilon)-neighbourhood is
$$ \mathcal{N}_\varepsilon(x_i) = \{ x_j \in \mathcal{X} \mid \lVert x_i - x_j \rVert \le \varepsilon \}. $$If (|\mathcal{N}_\varepsilon(x_i)| \ge \texttt{min_samples}|), the point is core. DBSCAN builds clusters by exploring density-reachable points; anything left unvisited becomes noise. With a spatial index the overall complexity is (O(n \log n)).
3. Python example #
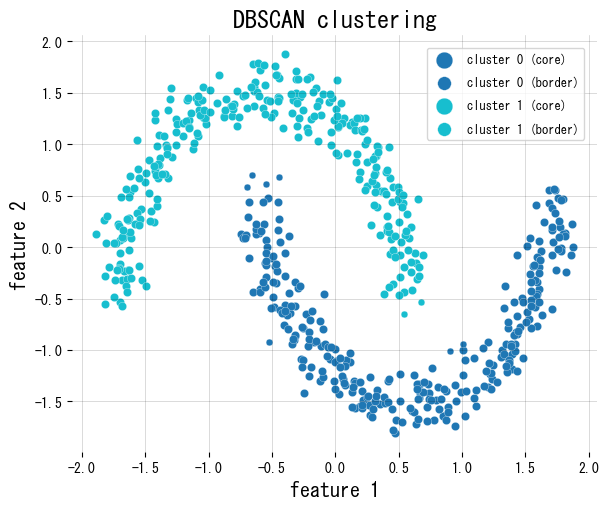
The snippet below uses scikit-learn’s DBSCAN on a two-moons dataset, colours core/border points differently, and reports how many samples are marked as noise.
| |
4. Practical tips #
- Plot the sorted distances to the k-th neighbour (k =
min_samples) to pickepswhere the curve exhibits an elbow. - Use pipelines so standardisation and clustering run together; otherwise distance-based decisions become meaningless.
- For very large datasets consider approximate nearest neighbours or switch to HDBSCAN, which extends DBSCAN to multi-density data and removes the need to tune
eps.
FAQ #
What is DBSCAN? #
DBSCAN (Density-Based Spatial Clustering of Applications with Noise) is an unsupervised clustering algorithm that groups points based on local density rather than distance to a centroid. It discovers clusters of arbitrary shape and automatically marks low-density points as noise (label -1), without requiring you to specify the number of clusters in advance.
What are the two main parameters of DBSCAN? #
eps(ε): the radius that defines a point’s neighbourhood. Two points are neighbours if their distance is ≤ ε.min_samples: the minimum number of neighbours within ε for a point to be considered a core point. Larger values create denser, smaller clusters and more noise.
A practical tuning recipe: fix min_samples to dimensionality + 1 (or higher for noisy data), then plot the sorted distance to the k-th nearest neighbour and pick ε at the “elbow” of the curve.
How does DBSCAN differ from k-means? #
| k-means | DBSCAN | |
|---|---|---|
| Number of clusters | Must be specified | Discovered automatically |
| Cluster shape | Convex (spherical) only | Arbitrary shape |
| Noise handling | All points assigned | Noise points get label -1 |
| Scalability | O(nk) per iteration | O(n log n) with index |
| Sensitive to | Cluster count k, outliers | eps, min_samples, scale |
Use k-means for compact, well-separated blobs; use DBSCAN when clusters have irregular shapes or you expect noise/outliers.
Why do I need to scale features before DBSCAN? #
DBSCAN uses a distance threshold eps to define neighbourhoods. If features have vastly different scales (e.g., age in years vs. income in thousands), the distance calculation will be dominated by the high-scale feature, making eps meaningless. Always apply StandardScaler or MinMaxScaler before fitting.
When should I use HDBSCAN instead of DBSCAN? #
Choose HDBSCAN when:
- Your dataset has clusters of varying density (DBSCAN’s single
epscannot handle this). - You want to avoid tuning
eps(HDBSCAN only needsmin_cluster_size). - You want a more stable solution across different datasets.
HDBSCAN builds a full density hierarchy and extracts the most persistent clusters, making it more robust than DBSCAN for real-world data.
5. References #
- Ester, M., Kriegel, H.-P., Sander, J., & Xu, X. (1996). A Density-Based Algorithm for Discovering Clusters in Large Spatial Databases with Noise. KDD.
- Schubert, E., Sander, J., Ester, M., Kriegel, H.-P., & Xu, X. (2017). DBSCAN Revisited, Revisited. ACM Transactions on Database Systems.
- scikit-learn developers. (2024). Clustering. https://scikit-learn.org/stable/modules/clustering.html