2.5.5
Gaussian Mixture Models (GMM)
- A Gaussian Mixture Model represents data as a weighted sum of multivariate normal components.
- It outputs a responsibility matrix that quantifies how strongly each component explains every sample.
- Parameters are estimated with the EM algorithm; covariance structures can be
full,tied,diag, orspherical. - Model selection typically combines information criteria (BIC/AIC) with multiple random initialisations for stability.
- k-means Clustering — understanding this concept first will make learning smoother
Intuition #
This method should be interpreted through its assumptions, data conditions, and how parameter choices affect generalization.
Detailed Explanation #
Mathematics #
The density of \(\mathbf{x}\) is
$$ p(\mathbf{x}) = \sum_{k=1}^{K} \pi_k \, \mathcal{N}(\mathbf{x} \mid \boldsymbol{\mu}_k, \boldsymbol{\Sigma}_k), $$with mixture weights \(\pi_k\) (non-negative and summing to 1). EM alternates:
- E-step: compute responsibilities \(\gamma_{ik}\). $$ \gamma_{ik} = \frac{\pi_k \, \mathcal{N}(\mathbf{x}_i \mid \boldsymbol{\mu}_k, \boldsymbol{\Sigma}_k)} {\sum_{j=1}^K \pi_j \, \mathcal{N}(\mathbf{x}_i \mid \boldsymbol{\mu}_j, \boldsymbol{\Sigma}_j)}. $$
- M-step: re-estimate \(\pi_k, \boldsymbol{\mu}_k, \boldsymbol{\Sigma}k\) using \(\gamma{ik}\) as weights.
The log-likelihood increases monotonically and converges to a local optimum.
Python walkthrough #
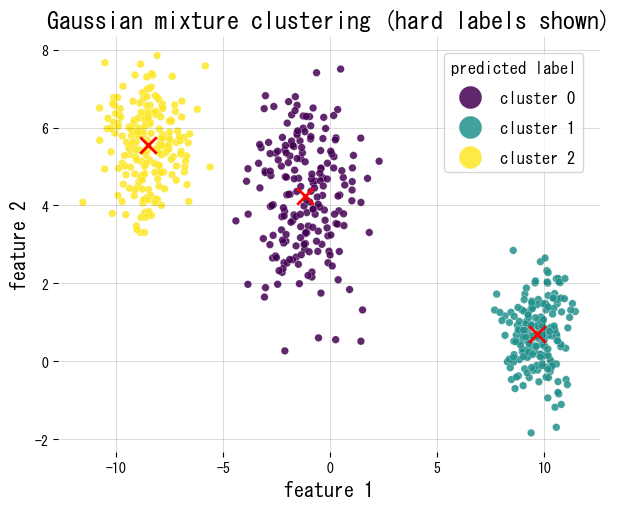
We fit a GMM to synthetic 2D blobs, plot the hard assignments, and report mixture weights and the responsibility matrix shape.
| |
FAQ #
What is a Gaussian Mixture Model? #
A Gaussian Mixture Model (GMM) is a probabilistic model that assumes data is generated from a mixture of \(K\) Gaussian (normal) distributions with unknown parameters. Each component has its own mean, covariance, and weight. Unlike k-means, which assigns each point to exactly one cluster, GMM produces soft assignments — a probability for each point belonging to each component.
How does a GMM differ from k-means? #
| k-means | GMM | |
|---|---|---|
| Assignment | Hard (one cluster per point) | Soft (probabilities to all clusters) |
| Cluster shape | Spherical (assumes equal variance) | Flexible (full, tied, diag, or spherical covariance) |
| Output | Cluster labels | Responsibility matrix + probabilities |
| Objective | Minimize within-cluster sum of squares | Maximize log-likelihood via EM |
GMM is better when clusters overlap, have different sizes, or non-spherical shapes. k-means is faster and simpler when clusters are well-separated and roughly equal.
How do you choose the number of components K? #
Fit GMMs for a range of \(K\) values and compare using information criteria:
- BIC (Bayesian Information Criterion): penalises model complexity more heavily; generally preferred for cluster selection.
- AIC (Akaike Information Criterion): less penalty for complexity; can overfit with many components.
Choose the \(K\) where BIC reaches a minimum or shows an elbow. Also run multiple initialisations (n_init) to avoid local optima.
What covariance types are available in scikit-learn? #
GaussianMixture supports four covariance structures:
full: each component has its own unconstrained covariance matrix — most flexible but most parameters.tied: all components share one covariance matrix — fewer parameters.diag: each component has a diagonal covariance (independent features, different scales per component).spherical: each component has a single variance value — closest to k-means.
Start with full when data is small; use diag or spherical when the feature dimension is high or data is limited.
What is the EM algorithm used in GMM? #
EM (Expectation-Maximisation) iterates two steps until convergence:
- E-step: given the current parameters, compute the responsibility \(\gamma_{ik}\) — the posterior probability that sample \(i\) belongs to component \(k\).
- M-step: update the mixture weights, means, and covariances using the responsibilities as soft weights.
The log-likelihood is guaranteed to increase (or stay the same) at each iteration, converging to a local optimum. Multiple restarts help find better solutions.
References #
- Bishop, C. M. (2006). Pattern Recognition and Machine Learning. Springer.
- Dempster, A. P., Laird, N. M., & Rubin, D. B. (1977). Maximum Likelihood from Incomplete Data via the EM Algorithm. Journal of the Royal Statistical Society, Series B.
- scikit-learn developers. (2024). Gaussian Mixture Models. https://scikit-learn.org/stable/modules/mixture.html