2.1.11
Weighted Least Squares (WLS)
Summary
- Weighted least squares assigns observation-specific weights so trustworthy measurements influence the fitted line more strongly.
- Multiplying squared errors by weights downplays high-variance observations and keeps the estimate close to reliable data.
- You can run WLS with scikit-learn’s
LinearRegressionby providingsample_weight. - Weights can stem from known variances, residual diagnostics, or domain knowledge; careful design is crucial.
Intuition #
This method should be interpreted through its assumptions, data conditions, and how parameter choices affect generalization.
Detailed Explanation #
Mathematical formulation #
With positive weights \(w_i\), minimize
$$ L(\boldsymbol\beta, b) = \sum_{i=1}^{n} w_i \left(y_i - (\boldsymbol\beta^\top \mathbf{x}_i + b)\right)^2. $$The optimal choice \(w_i \propto 1/\sigma_i^2\) (inverse variance) gives more influence to precise observations.
Experiments with Python #
We compare OLS and WLS on data whose noise level differs across regions.
| |
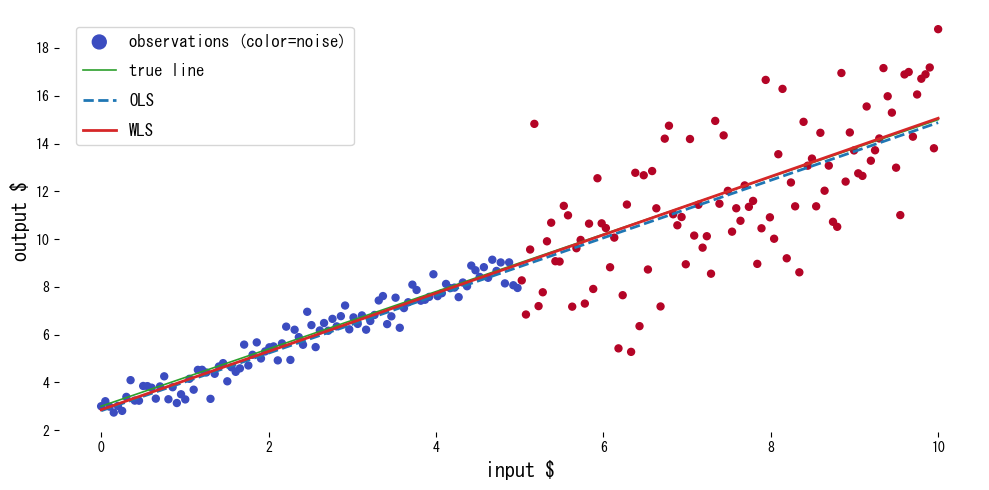
Reading the results #
- Weighting draws the fit toward the low-noise region, producing estimates close to the true line.
- OLS is skewed by the noisy region and underestimates the slope.
- Performance hinges on choosing appropriate weights; diagnostics and domain intuition matter.
References #
- Carroll, R. J., & Ruppert, D. (1988). Transformation and Weighting in Regression. Chapman & Hall.
- Seber, G. A. F., & Lee, A. J. (2012). Linear Regression Analysis (2nd ed.). Wiley.